
Automated microscopes that adapt to each sample’s quirks can capture elusive biological phenomena at high resolution.

Physicist Robert Prevedel builds microscopes that can quickly measure neuron activity while adapting to movements in the live mouse brain. Credit: EMBL
A mouse’s heart beats roughly 600 times each minute. With every beat, blood pumping through vessels jiggles the brain and other organs. That motion doesn’t trouble the mouse, but it does pose a challenge for physicist Robert Prevedel.
Prevedel designs and builds microscopes to solve research problems at the European Molecular Biology Laboratory in Heidelberg, Germany. In his case, the issue is how to capture neural activity deep inside the brain when the organ itself is moving. “The deeper we image, the more aberrations we accumulate,” he says. “So, we have to build our microscope to quickly measure and adapt.”
Typically, microscopes struggle to peer deeper than about one millimetre into tissues. Beyond that, light bouncing off intracellular structures creates distortions, blurring the images — even when the sample is not moving to a heartbeat.
In 2021, Prevedel and his colleagues engineered a microscope that combines an approach called three-photon fluorescence imaging, which is used to probe inside tissues, with an adaptive optical system. The latter strategy was first developed in astronomy, and directs light using mirrors and lenses made of deformable membranes instead of rigid optical materials. Software tools quickly alter the membranes’ shapes in response to variations in the sample, bending light in specific ways to peek through surface structures. Rather than needing a human operator, this ‘sample-adaptive’ approach relies on the microscope itself adjusting the optics in real time to continually produce high-quality images.
Prevedel’s microscope had to go one step further to compensate for the vibrations of the animal’s pumping heart. By timing photos to the heartbeat, the team was able to image cells nearly 1.5 millimetres beneath the brain’s surface in a region called the hippocampus, and 0.5 mm deeper than previous efforts1. “It was really exciting to see that it worked much farther down than we were expecting,” Prevedel says.
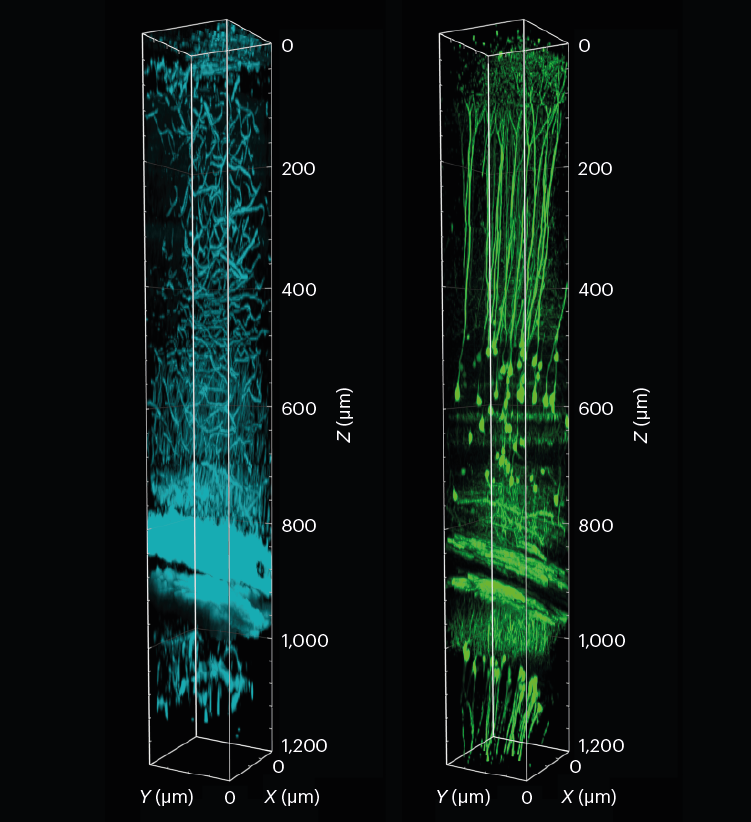
A 3D reconstruction of a section of the mouse visual cortex and hippocampus, showing tissue structure (cyan) and neurons (green). The sample was imaged to a depth of 1.2 mm using three-photon fluorescence microscopy with an adaptive optical system.L. Streich et al./Nature Methods (CC BY 4.0)
Prevedel’s microscope is just one of a suite of smart microscopes that rely on adaptive optical systems such as deformable membranes, coupled with machine-learning approaches, to peer deeper into tissues than ever before, and to zoom in at crucial moments to capture fleeting moments in the life of a cell.
Often involving custom-built hardware and software components that can make them inaccessible to the broader biological community, these efforts are producing tools that do more than just produce clear images. They’re also gentler on living systems, allowing researchers to observe prolonged processes without damaging their samples, thus expanding the reach of microscopy.
“If all your research involves looking at cells on slides, you don’t need adaptive microscopy,” says Martin Booth, an optical-systems engineer at the University of Oxford, UK. “But if you’re going tens or hundreds of micrometres deep into living specimens, that’s when this becomes really useful.”
For Kate McDole, now a developmental biologist at the MRC Laboratory of Molecular Biology in Cambridge, UK, the key consideration in designing her adaptive microscope was sample size.
As a postdoctoral fellow at the Howard Hughes Medical Institute’s Janelia Research Campus in Ashburn, Virginia, McDole and her colleagues developed a microscope to study how clumps of cells form complex tissues as the mouse embryo develops2. Led by developmental biologist Philipp Keller, the team wanted to image the developing embryo over three days, during which its diameter grows from about 200 micrometres to nearly 3 millimetres. Anchored on one end, the embryo, McDole says, is “free to sway in the breeze — it moves”, and its density and other optical properties change over time. “The microscope needed to keep up with all of that,” McDole says.
She and her colleagues started with a technique known as simultaneous multiview light-sheet microscopy, which Keller’s team had developed to track cell movement in developing fruit-fly embryos. To adapt it for the mouse, they altered the optical design and built software to control such factors as the angle of the light source and the positions of optical elements. The software was able to gauge the data quality of the images as they were collected, and could tweak these factors to optimize images throughout the experiment.
The microscope was also automated to detect the growing embryo’s position in the sample chamber and keep it centred in the field of view, adjusting the distances to ensure consistent image quality. “A lot of biological specimens like to roam around, so you need to keep them centred,” McDole says.
McDole and her team used this system to observe mouse embryo development over a 48-hour period, imaging the embryonic heart and other developing organs at single-cell resolution. They have subsequently imaged brain organoids for up to two weeks. “That’s kind of the future of autonomous microscopy — letting the microscope decide when and where and how to act on specific events,” McDole says. “You can teach the microscope: ‘It’s going to be 3:00 a.m. when this cell division happens, and I want you to zoom in and do something and then go back to normal imaging.’”
The Keller group has released schematics for building such an instrument, and the software is freely available online (see go.nature.com/3huukh9). Researchers who already use a light-sheet microscope can expect to spend about US$30,000 to add these capabilities to their system, Keller says. “It’s a pretty small extra investment” to achieve deep-tissue, single-cell resolution, he notes.
Smart microscopes are also being developed to image smaller structures. Physicist Ilaria Testa at the KTH Royal Institute of Technology in Stockholm, for instance, built one to observe subcellular vesicles as they release calcium at neural synapses when nerve cells fire — a key step in signal transmission. “These are rare events and it’s not always easy to capture them,” she says.
One option was to image a specimen continually in the hope of capturing the moment. But vesicle-release events are transient and the structures too small for standard microscopy. Super-resolution imaging can reveal more detail, but it requires high-intensity light sources that can be used only briefly before the sample is damaged. The team tried various time-lapse approaches, in which images are captured at regular intervals. But that, Testa says, was like watching a soccer match and missing a goal because you are looking elsewhere at the crucial moment. “It was a bit frustrating,” she says. “The resolution would have been fine if we only knew where to look.”
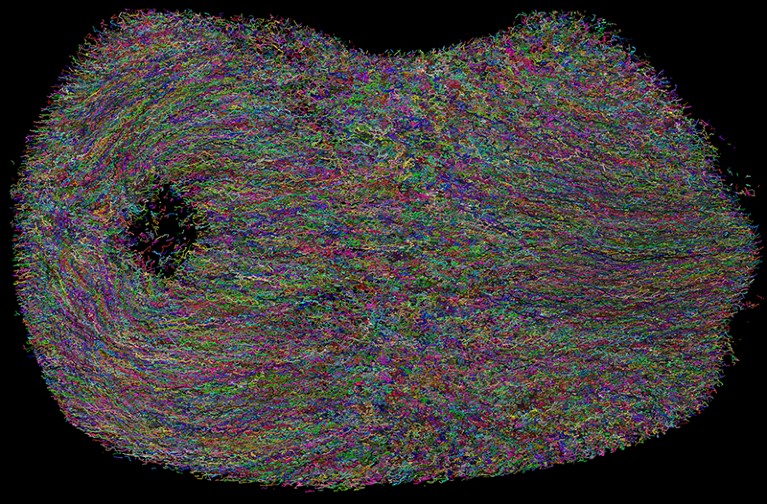
Individual cell tracks in a developing mouse embryo, imaged over 48 hours using adaptive light-sheet microscopy and a deep-learning algorithm for automated tracking.Credit: Kate McDole
To help them keep their eyes on the ball, Testa and her team combined two microscopy approaches: a fluorescent wide-field microscope and a form of super-resolution microscopy known as stimulated emission depletion (STED). They developed a software system to control these microscopy modes: when the software detected a change in fluorescence, the system would switch automatically to the higher-resolution STED mode. This allowed the team to capture — with nanometre precision — how cells reorganize their synaptic vesicles after releasing calcium3. “We’re basically guiding the acquisition of the image in a smarter way,” Testa says.
Testa and her team took about two years to build the microscope, she says, using mostly custom 3D-printed hardware. The only commercial component was the upper part that includes the eyepiece, known as the microscope head. The set-up cost about $200,000: most expensive were the lasers, which cost some $60,000. But the result is an automated system that works more efficiently than a researcher trying to switch between microscopes, Testa says. “We’re actually delegating something that the computer can do much better than a human.” The set-up also reduces the effort required to home in on the fleeting moments of biology that really matter, Prevedel adds. “The microscope just acquires the few seconds that you really care about.”
Testa’s software is available on GitHub and her team has published the hardware schematics3, so the microscope could — in principle — be used by anyone. But it requires a super-resolution microscope, which not all labs are able to access, particularly in resource-poor regions of the world, she points out.
For now, researchers using sample-adaptive approaches tend to collaborate with colleagues who can build or modify custom microscopes. But when it comes to DIY projects, software and instrument tweaks are often required, notes Booth. The Keller team spent months identifying the best metrics and computational strategies to use in their adaptive microscope, McDole says. Even so, the equipment is often constantly changing, and “users require significantly more training than on your standard commercial microscope”, she says.
Microscopy: hello, adaptive optics
Biologists wanting to team up with engineers to access custom microscopes might only be able to do so at large, well-funded universities and research centres. Cell biologist Ricardo Henriques was keenly aware of this issue because of his experiences working in the United Kingdom and South Africa during his graduate studies.
“Almost everything that we’ve done in terms of creating technology has been designed to be accessible to everyone,” says Henriques, who now leads a research group at the Gulbenkian Institute of Science in Lisbon in his native Portugal. “We are particularly sensitive to the fact that there’s a big difference between the global south and the global north when it comes to accessing these resources.”
In 2016, Henriques and his team devised a computational method called super-resolution radial fluctuations (SRRF) to extend super-resolution microscopy to conventional fluorescence microscopes, which are generally widely accessible4. The method now relies on a machine-learning approach that adapts the microscope to acquire images at high speeds and then parse the data to identify sample-specific blips in fluorescence. Users capture 100 images of their sample at high speeds to minimize chemical damage to fluorescent labels, known as bleaching, as well as to reduce blurring from movements. The data are then fed into an algorithm that looks for variations in fluorescence.
The fluorescent molecules, or fluorophores, that are used to label cellular structures fluctuate in brightness depending on their location and other sample properties. “Every time [the algorithm] picks up a fluctuation, it’s able to pinpoint the exact centre of the fluctuation and if it’s coming from a fluorophore or not,” Henriques explains. “So, it generates something that’s crisper than the original.”
Lattice light-sheet microscopy gets an AO upgrade
Microscopist Adán Guerrero at the National Autonomous University of Mexico’s National Advanced Microscopy Laboratory in Cuernavaca was one of the first researchers to adopt the SRRF method, choosing it to explore the subcellular location of a bacterial protein5. Guerrero’s institute had super-resolution microscopes at the time, but users often had to wait weeks to use them. Because Henriques and his team had released their SRRF tool as a plug-in for the image-processing program ImageJ, it could be used with the institute’s readily available microscopes, making the tool especially practical, Guerrero says. The plug-in element was also valuable, he adds, because his users were uncomfortable doing their own coding. “There were other, similar technologies published earlier or around the same time, but the speed of getting data and the plug-in made SRRF more accessible.”
The possibility of obtaining higher-quality images without the need for expensive instrumentation has made a huge difference to his research community, Guerrero adds. But users must still cross-validate their images with other techniques, such as biochemical studies or electron microscopy, to avoid mistakes, warns Caron Jacobs, an imaging scientist at the University of Cape Town in South Africa.
For instance, when Jacobs was a graduate student in Henriques’ lab, she tried to image how HIV engages with protein receptors on the surfaces of the immune cells that the virus infects, called T cells. After using SRRF, Jacobs and her colleagues saw a distinct “honeycomb-like lattice” — but the protein receptors, the team knew, were actually evenly distributed. “As we saw the honeycomb pattern, we knew it was an artefact,” Jacobs says. “But if you don’t know the underlying biology, there’s a danger of thinking it’s real.”
The microscope makers putting ever-larger biological samples under the spotlight
Henriques’ team has developed another computational tool called SQUIRREL (super-resolution quantitative image rating and reporting of error locations) that allows users to check their images for such artefacts6. But similar problems can occur with any computationally intense microscopy technique, says Jacobs, who now helps biologists to image a wide range of samples. When students approach her for advice on super-resolution or other microscopy techniques, Jacobs encourages them to think about the data they want. “If you drill down to the scientific question, nine times out of ten they don’t need either [approach] — just good wide-field microscopy and sample-preparation strategies.”
